Since the late 19th century, research for human embryology relied on the morphological observations of human embryonic samples, including observation of the gross view using histological sectioning. For understanding the complex morphogenetic processes that occur during embryonic development, serial histological sections were utilized not only for histological two-dimensional (2D) observation but also for designing three-dimensional (3D) plaster or wax models (O’Rahilly and Müller, 1987; Morgan, 2004). Thus, studies of human embryology were the works of a team of scientists and technical specialists such as illustrators and modelers. The historical models of Ziegler, Blechshmidt, and Heard have been world widely famous and indispensable in research as well as teaching (Morgan, 2004).
For an administrator of human embryo collections, the preservation and maintenance of the collections is an important issue. Human embryo collections are usually held as fixed embryos of the whole body or sectioned tissue as well as thinly sectioned tissue on glass slides with staining for histological observations. Most samples were collected more than 30 years ago. The digitization of the histological glass slides is one solution for preserving the collections and for decreasing the maintenance cost. The Digital Embryology Consortium, an international partnership, was established in 2014 to digitize, preserve, and disseminate the major embryology histological collections for researchers (see https://embryology.med.unsw.edu.au/embryology/index.php/Digital_Embryology_Consortium_-_Information).
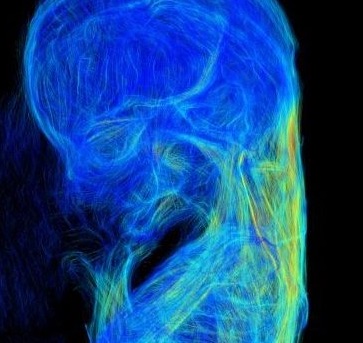
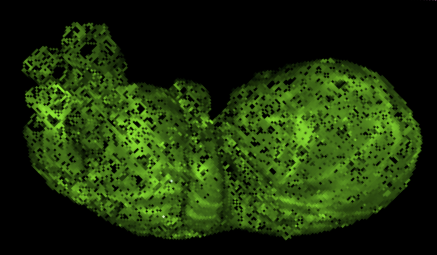
Digitization of whole embryo with the emergence of imaging systems is also on-going in several collections. Today, whole embryos are imaged three-dimensionally with various imaging modalities such as magnetic resonance imaging (MRI), phase-contrast X-ray computed tomography (PCT), and optical projection tomography (OPT). These imaging modalities are non-invasive and non-destructive methods that avoid destroying the precious human embryos. Digitized embryos as well as digitized serial histological sections and processed 3D reconstructed objects have been made available on several web sites, including the multi-dimensional human embryo (Smith,1999) at http://embryo.oad.umich.edu/index.html; the virtual human embryo, which is a digital image database of serially sectioned human embryos from the Carnegie Collection at http://virtualhumanembryo.lsuhsc.edu; the visible embryo at http://www.visembryo.com/baby/index.html; the virtual human embryo (Gasser et al., 2014) at http://www.ehd.org/virtual-human-embryo/; the Kyoto Human Embryo Visualization Project at http://atlas.cac.med.kyoto-u.ac.jp; The Human Developmental Biology Resource (Kerwin et al., 2010) , and the Atlas of Human Embryos, which is an interactive 3D atlas using episcopic fluorescence image capturing (EFIC) and MRI (Yamada et al., 2010).
The digitized materials are primarily used for education, as they are attractive materials for students and researchers who could be apprehensive. For research, they are subjected to use as database and references. We believe such digitized datasets contain a huge amount of information and have the potential to serve as the main component for researches, even though current usage of these datasets is still limited to support materials. Recently, we started 3D analysis of the human embryonic period using digitized datasets obtained from the Kyoto Collection. Indeed, the digital datasets play the primary role in this research. The features of our current research and prospects for the future use of these datasets will be described.